How will the virtual meeting work?
For detailed instructions, please carefully read the MLHC 2020 Attendee Guide.
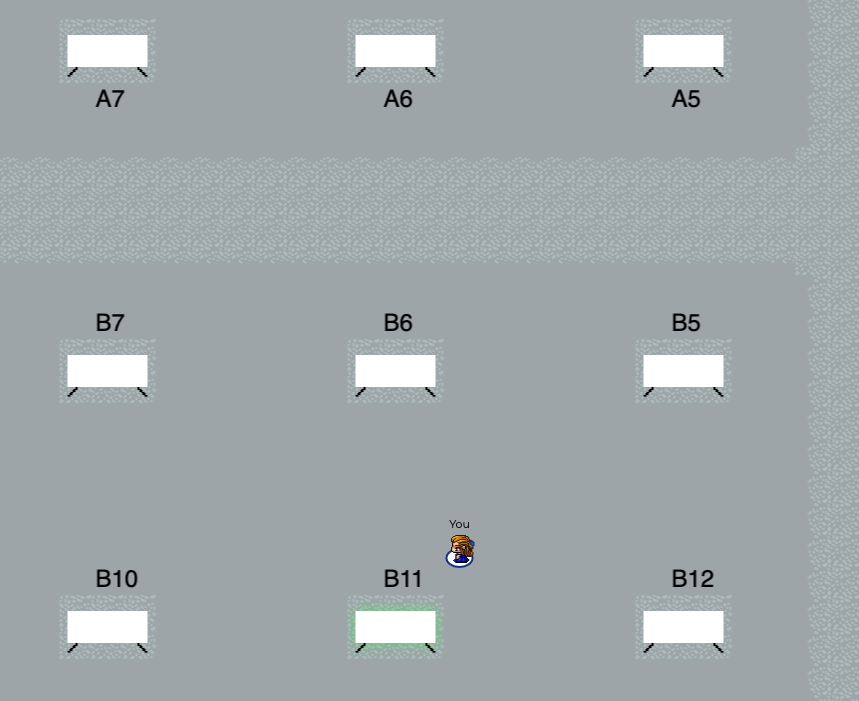
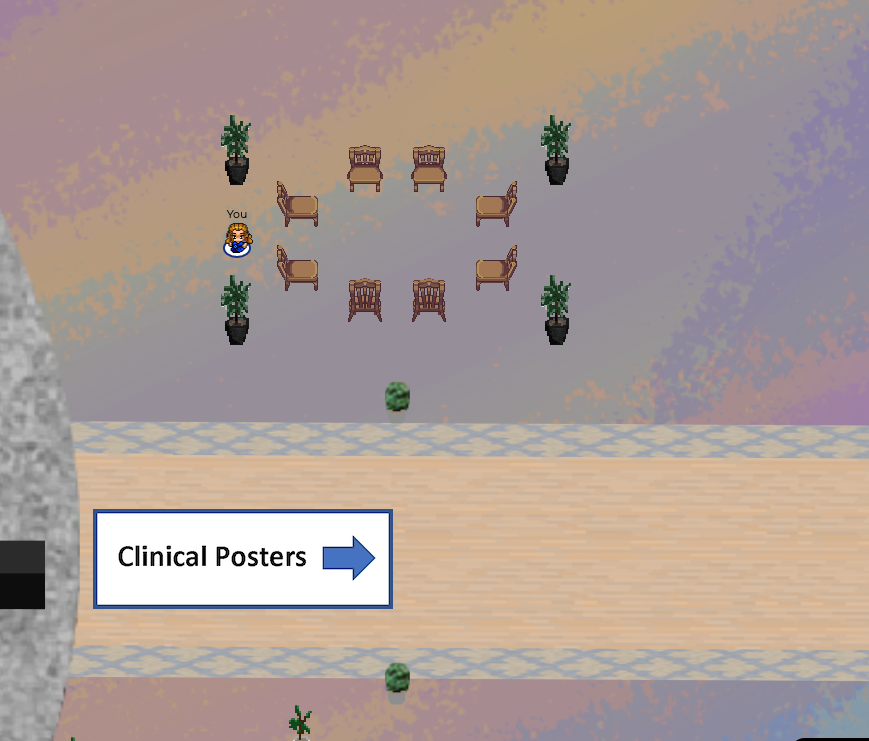
All invited talks have been prerecorded and are available on our MLHC YouTube channel, all accepted papers and abstracts are associated with a prerecorded spotlight presentation hosted on our YouTube channel (Posters A, Posters B, Clinical Abstracts). Most of Aug. 7th and 8th will be spent in our virtual 2-dimensional MLHC world created by gather.town. You’ll be able to walk among the posters, interact with poster presenters, and network with other conference attendees (see screenshot below). Live Q&A sessions will be held in the ‘main auditorium’ of the virtual world through GoToWebinar. Registered participants will receive additional instructions in the days leading up to the meeting. We look forward to seeing you in 2D!
2020 Main Conference Agenda
Friday, August 7th, 2020, Virtual (all times are EDT)
____________________________________________________________________________
Please note that all talks (invited and submitted) are available on our YouTube channel and can be viewed at any time. The schedule below only pertains to interactive portions of the meeting including moderated discussion with invited speakers and the poster sessions. All times are in EDT.
-----Session 1 -----
Moderated Discussion/Q&A with Invited Speakers [GoToWebinar]
Moderator: Finale Doshi-Velez, PhD John L. Loeb Associate Professor in Computer Science, Harvard University
10:30 - 10:50 Robert Califf, MD, Head of Medical Strategy and Policy for Verily Life Sciences and Google Health
Title: Opportunities in a Digital Clinical World - Before and After the Pandemic
11:00 - 11:20 Emma Brunskill, PhD, Assistant Professor, School of Computer Science, Stanford University
Title: Learning from Little Data to Robustly Make Good Decisions
---Poster Session A & Breakouts--- [gather.town]
11:30 - 12:00 Breakout Sessions:
Breakout Room 1: From Clinic to Community: ML and Social Determinants of Health, with Dan Lizotte: Do you use social determinants of health information (e.g. gender, socioeconomic status, racial identity) in your models? Why or why not? How can we use this kind of information in a responsible way?
Breakout Room 2: From Predictions to Decisions: How to make ML4HC Actionable, with Zachary Lipton: Despite the surge of activity in applications of modern ML techniques to healthcare data and public excitement about revolutionizing care, it's often unclear how the predictions, representations, etc. learned by ML algorithms can or should be incorporated into treatment decisions. In this breakout session, we'll discuss the most promising areas to have real impact on clinical care, and the technical challenges that must be overcome to achieve wider (positive) impact.
Breakout Room 3: Using Causal Inference and Transfer Learning for Practical Decision Making in Heterogeneous Populations, with Sonali Parbhoo: How we can help address causal queries in more practical ways e.g. through combining observational and interventional data or improving existing benchmarks for causal inference, as well as discussing the intersection between RL and causal inference? I also think it would be interesting to discuss ways in which one could transfer the knowledge gained from data in well-resourced countries to those with less resources to bring about practical improvements in these communities (eg. via learning better representations).
Breakout Room 4: Learning health from Time Series: The Time is now! with Luca Foschini: Data from wearable devices, remote monitoring and telehealth system are being produced at unprecedented pace, and when coupled with symptom tracking and a data infrastructure that guarantees privacy they can help understand health and disease outside the clinic walls. Let's discuss opportunities of ML in continually learning health from time series from millions of people: what are meaningful ML tasks and what models tend to perform well in these regimes? Can self-supervised learning help across the board? What are the opportunities for causal inference in these settings?
Breakout Room 5: What are Suitable Benchmark Tasks for ML in Healthcare? with Jason Fries: Shared benchmarks drive algorithm development in machine learning. In NLP, multi-task datasets such as SuperGLUE assess performance across a variety of tasks. What shared tasks would make good benchmarks for ML in healthcare?
Breakout Room 6: ML/Health Research and Opportunities in Industry with Emily Fox: What is it like to do ML/health-related research in industry? What are the challenges? What are some of the opportunities?
Breakout Room 7: Sensitivity and Robustness of Machine Learning Analyses with Soumya Ghosh: Measuring sensitivity and robustness of ML methods to perturbations in training data and/or modeling assumptions is essential for healthcare applications. Does your ML workflow include sensitivity analysis? What techniques do you rely on for quantifying sensitivity? Does your favorite technique account for temporal correlations typical in healthcare data? We will discuss these issues and highlight common tools and compute efficient approximations for such analysis, in this breakout session.
11:30 - 13:30 Papers Research Track Posters A [gather.town]
-----Session 2 -----
Moderated Discussion/Q&A with Invited Speakers [GoToWebinar]
Moderator: Byron Wallace, PhD Assistant Professor of Computer Science, Northeastern University
13:30 - 13:50 Besmira Nushi, PhD, Senior Researcher in the Adaptive Systems and Interaction, Microsoft Research AI
Title: The Unpaved Path of Deploying Reliable and Human-Centered Machine Learning Systems
Abstract: As Machine Learning systems are increasingly becoming part of user-facing applications, their reliability and robustness are key to building and maintaining trust with users, especially for high-stake domains such as healthcare. While advances in learning are continuously improving model performance in expectation and in isolation, there is an emergent need for identifying, understanding, and mitigating cases where models may fail in unexpected ways and therefore break human trust or dependencies with other larger software ecosystems. Current development infrastructures and methodologies often designed with traditional software in mind, still provide very little support to enable practitioners debug and troubleshoot systems over time. This discussion will look at such problems from two different stakeholder lenses: machine learning practitioners and end user decision makers. From a practitioner perspective, it will summarize some of the current gaps in tooling for responsible ML development and evaluation, and present ongoing work that can enable in-depth error analysis and careful model versioning. Next, from an end user perspective it will propose rethinking the optimization of machine learning models such that it takes into consideration human-centered properties of human-machine collaboration and partnership. While both these lenses pose both research and engineering practices, they also require close collaboration with domain experts who are using machine learning in the open field to ensure that deployed systems meet real-world expectations.
14:00 - 14:20 Ziad Obermeyer, MD, MPhil, Acting Associate Professor of Health Policy and Management, School of Public Health, UC Berkeley
Title: Algorithms are as good as their labels
---Poster Session B--- [gather.town]
14:30 - 16:30 Paper Research Track Posters B [gather.town]
Saturday August 8, 2020, Virtual
____________________________________________________________________________
-----Session 3 -----
Moderated Discussion/Q&A with Invited Speakers [GoToWebinar]
Moderator: James Fackler, MD, Associate Professor of Anesthesiology and Critical Care Medicine and Pediatrics, Johns Hopkins
10:30 - 10:50 Madeleine Clare Elish, PhD, Program Director and co-founder of the AI on the Ground Initiative, Data & Society
Title: Repairing Innovation: The Labor of Integrating New Technologies
11:00 - 11:20 David Sontag, PhD, Associate Professor of Electrical Engineering and Computer Science, MIT
Title: Machine Learning to Guide Treatment Suggestions
---Poster Session C & Breakouts--- [gather.town]
11:30 - 12:00 Breakout Sessions:
Breakout Room 1: Causal inference in practice, with Uri Shalit: We will discuss thoughts, experiences and questions about integrating causal inference methods into real-world medical systems.
Breakout Room 2: Practical Applications of Reinforcement Learning in Healthcare, with Yuan Luo: Large healthcare chains such as Northwestern Medicine has curated clinical, genetic and imaging data of >8 million patients, along with their interventions. This could be a rich oil field for RL to drill in, but so far successful applications seem less often than desired. This breakout session can serve the purpose of introducing people interested in RL who may be looking for either data or suitable methods.
Breakout Room 3: Fusion of Multimodal Health Data, with Ina Fiterau: Does your healthcare application involve data of varied types, such as time series (e.g., vital signs, activity data) and images (e.g., xRays/MRIs), perhaps in conjunction with structured tables? Do you often find yourself having to train separate models to extract representations from unstructured data? Or perhaps excluding specific data because the format is difficult to work with? Well, in this breakout we'll discuss different techniques for nontrivially merging data types and mining your messy multimodal data for all its worth, all to the benefit of health.
Breakout Room 4: Moving from Academia to Industry in Health Research, with Katherine Heller: I will talk about the effects on health research that a move from academia to industry (tech) has. What are the differences in the work that goes on or what can be accomplished?
Breakout Room 5: NLP for Healthcare, with Tristan Naumann: Much information recorded in a clinical encounter is located exclusively in provider narrative notes, which makes them indispensable for supplementing structured clinical data in order to better understand patient state and care provided. Join us in discussing: opportunities afforded by NLP in healthcare, common NLP tasks in healthcare, NLP tools (tell your cTAKES story!), medical ontologies, and more!
Breakout Room 6: Privacy in MLHC, with Lovedeep Gondara: We will discuss the use of differential privacy to create ML models for healthcare, including predictive and generative; addressing the privacy-utility bottleneck.
Breakout Room 7: Preventing Machine Learning Models from Biasing Future Data, with George Adam: We will explore how ML models interacting with clinicians can have a larger than intended effect on clinician decision making. We will discuss how to prevent ML models from reinforcing their prediction bias when they are regularly updated, and are able influence future labels via their predictions.
11:30 - 13:30 Clinical Track Posters [gather.town]
-----Session 4 -----
Moderated Discussion/Q&A with Invited Speakers [GoToWebinar]
Moderator: Michael Sjoding, MD, Assistant Professor of Critical Care Medicine, University of Michigan
13:30 - 13:50 Nicholson Price, PhD, JD, Assistant Professor, Michigan Law, University of Michigan
Title: Legal Regimes and the Spectrum of Medical AI/ML
Abstract: Biomedical technology is profoundly shaped by three interacting legal regimes: FDA regulation, the patent system, and insurance reimbursement. For a small fraction of medical AI--commercially developed, FDA-cleared point-of-care systems--these regimes are present in nonstandard but still highly salient ways. But for a very large fraction of medical AI, including most user-developed AI and most AI used further from the point of care, these regimes are much less dominant and operate in different ways, with implications for what gets developed, who does the developing, and the efficacy and fairness of the resulting systems.
14:00 - 14:20 Leora Horwitz, MD, MHS, Associate Professor, Department of Medicine, NYU Langone Health
Title: A clinician's perspective on machine learning in healthcare
-----Session 5 -----
Panel Discussion [GoToWebinar]
Moderator: Rajesh Ranganath, PhD, Assistant Professor of Computer Science and Data Science, NYU
15:00 - 16:00 Heterogeneous Treatment Effect Estimation
Panelists:
Issa Dahabreh, ScD, Associate Professor of Health Services, Policy and Practice, Associate Professor of Epidemiology
David Kent, MD, CM, MS Professor of Medicine, Neurology and Clinical and Translational Science
Suchi Saria, PhD, John C. Malone Associate Professor of Computer Science at the Whiting School of Engineering and of Statistics and Health Policy at the Bloomberg School of Public Health
David Sontag, PhD, Associate Professor of Electrical Engineering and Computer Science, MIT
Feedback Session [Zoom]
16:00 - 16:30 Open feedback session with the MLHC Organizers to discuss ways to improve the conference in the future.